In recent years, life sciences have witnessed the advent of new high-throughput sequencing technologies. These generate vast quantities of data that cannot be processed manually. The CRA-W has therefore acquired human and technological resources in bioinformatics.
Bioinformatics can be applied to various areas of life sciences, including genomics (the study of DNA), transcriptomics (the study of expressed genetic information) and proteomics (the study of proteins) to name but a few. However, the basic principle in these different approaches is quite similar: starting from complex raw data and applying a series of commands to process them (cleaning, sequence alignment, taxonomic assignment, etc.), thus enabling conclusions to be drawn. Bioinformaticians are regularly required to write scripts, i.e. a set of commands dedicated to a section of the bioinformatics pipeline. These scripts are written in various programming languages (e.g. R, bash, python, etc.) and it is not uncommon for scripts written in different languages to be used within the same pipeline.
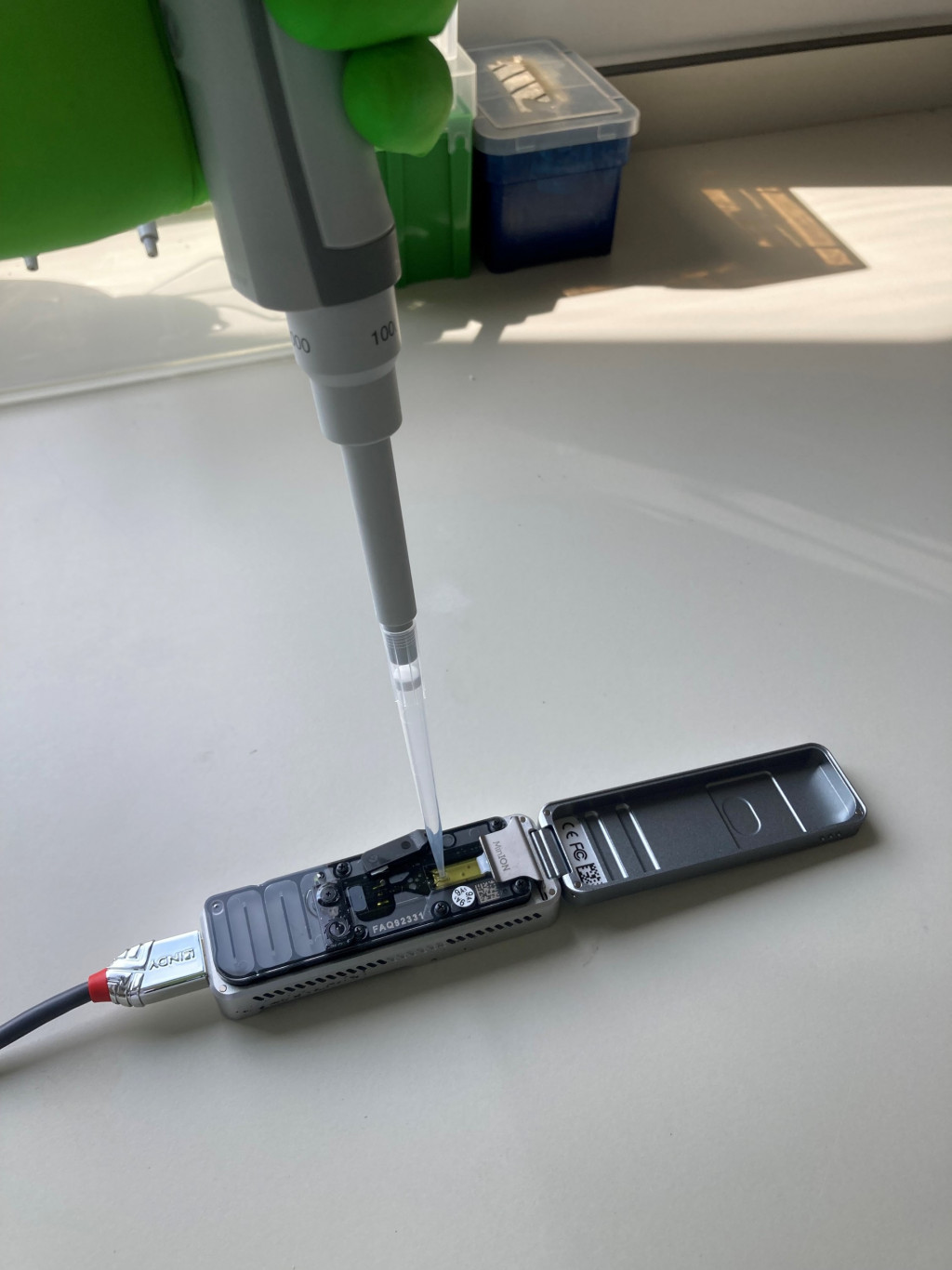
One of the most important applications associated with bioinformatics is the study of samples by metabarcoding. Through the massive amplification of certain genomic sequences of interest, bioinformatics can be used to determine the taxonomic composition of samples that may be very different: fungal spores collected in the air, pollen, soil, plants, faeces, food products and many others. The number of possibilities has expanded with the arrival of a new generation of sequencers (e.g. sequencing by nanopores on a portable MinION-type device), which means that much longer fragments can be sequenced. These possibilities include better identification of the organisms present in a sample, or the sequencing of complete genomes. Several CRA-W teams have recently acquired this type of sequencers.
A number of high-throughput sequencing activities are taking place in several CRA-W units to study a wide range of organisms: plants, animals (including insects), bacteria, fungi, viruses, etc.
These bioinformatics skills are offered to various CRA-W units or other research institutions to meet the demands associated with identifying the species present in a sample or characterising microbial communities. The CRA-W thus hopes to position itself as a leading player in the analysis of data obtained by high-throughput sequencing in connection with agronomic and environmental research.
Photo caption: New generation portable sequencer acquired by several CRA-W teams.